NOTE: The information on this post may be outdated. We recommend looking into the up-to-date documentation at trac (internal) and ReadTheDocs.
To add data to any table, there are two easy steps. As our working example, we’ll add some new objects to the SOURCES table.
Step 1: Create an ascii file of the data
First, you must choose a delimiter, which is just the character that will break up the data into columns. I recommend a pipe ‘|’ character since they don’t normally appear in text. This is better than a comma since some data fields may have comma-separated values.
Put the data to be added in an ascii file with the following formatting:
- The first line must be the |-separated column names to insert/update, e.g.
ra|dec|publication_id. Note that the column names in the ascii file need not be in the same order as the table. Also, only the column names that match will be added and non-matching or missing column names will be ignored, e.g.spectral_type|ra|publication_id|decwill ignore thespectral_typevalues as this is not a column in the SOURCES table and input the other columns in the correct places. - If a record (i.e. a line in your ascii file) has no value for a particular column, type nothing. E.g. for the given column names
ra|dec|publication_id|comments, a record with nopublication_idshould read34.567|12.834||This object is my favorite!.
Step 2: Add the data to the specified table
To add the data to the table (in our example, the SOURCES table), import astrodbkit and initialize the .db file. Then run the add_data() method with the path to the ascii file as the first argument and the table to add the data to as the second argument. Be sure to specify your delimiter with delim='|'. Here’s what that looks like:
from astrodbkit import astrodb
db = astrodb.Database('/path/to/the/database/file.db')
db.add_data('/path/to/the/upload/file.csv', 'sources', delim='|')
That’s it!




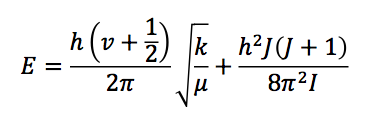
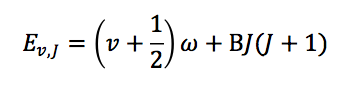
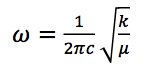 and
and