Over the course of this semester, I have been utilizing the program Sherpa, written in Python, to model Potassium (K) absorption lines and make three specific measurements. Specifically, these measurements are the equivalent width, full width half maximum, and line-to-continuum flux ratio. In this entry, I will show the commands I utilized to make these measurements.
When I use Sherpa, I import it as a python module as such:
>>> from sherpa.astro.ui import * # All Sherpa functions are found here
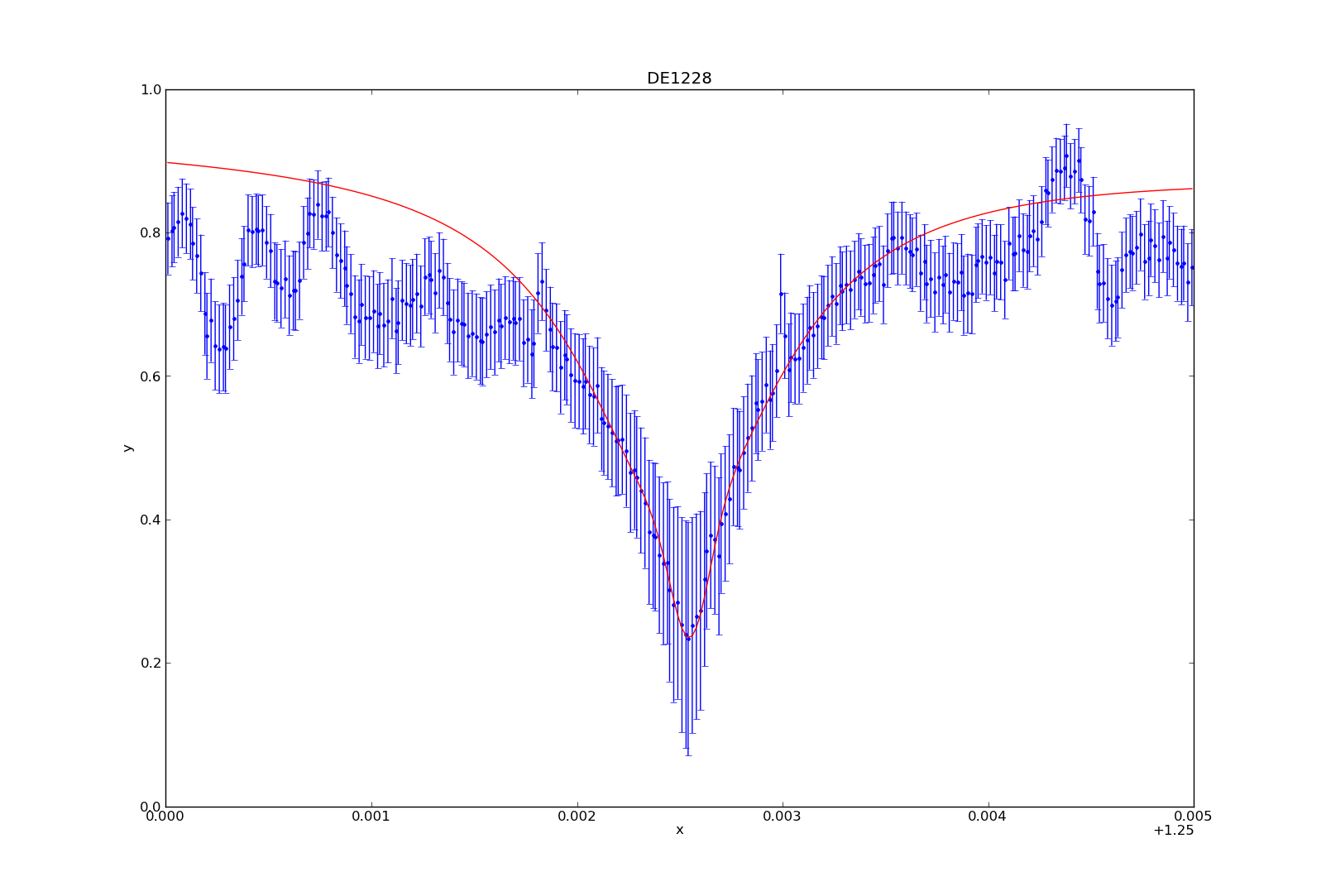
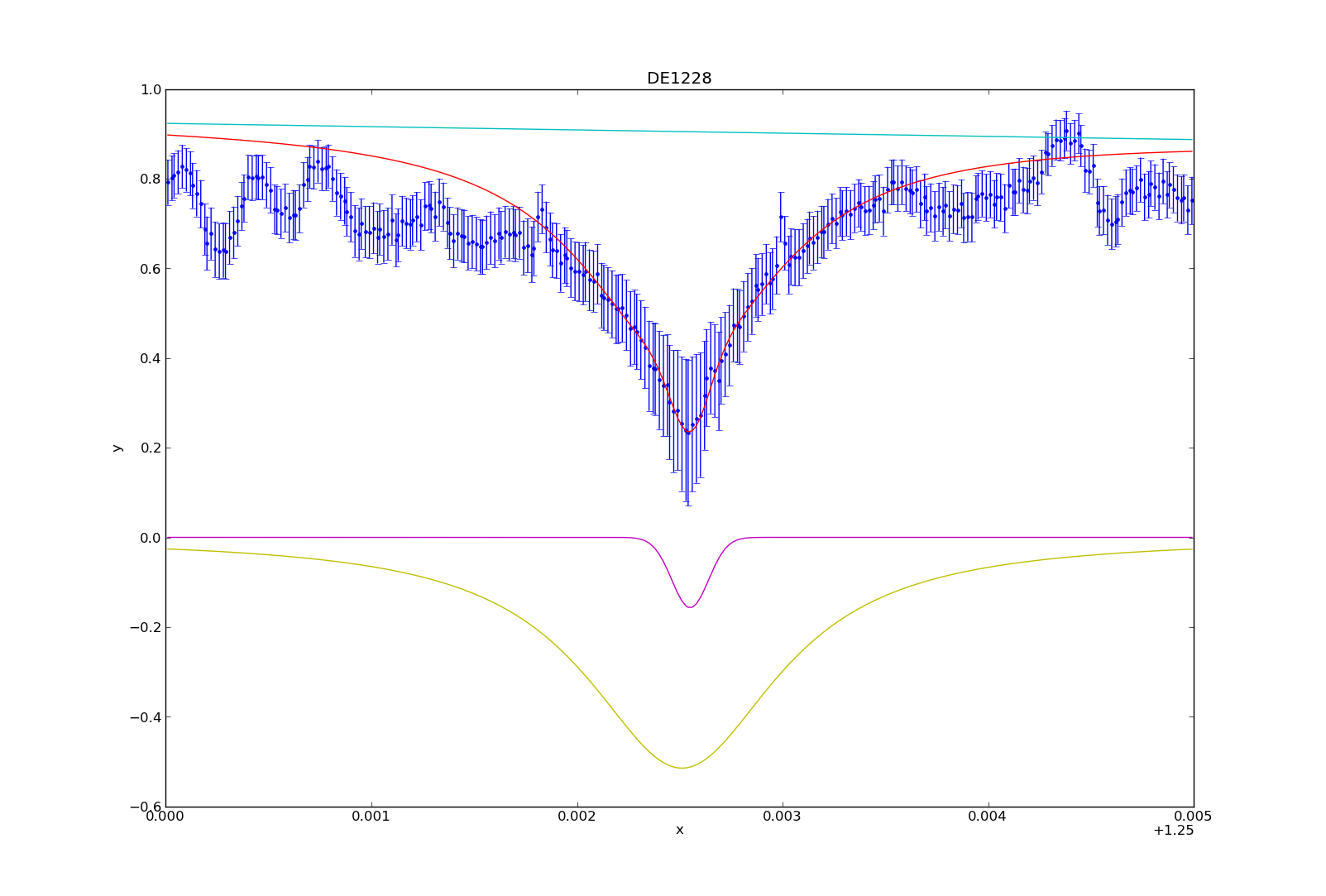
If you followed my post on modeling with Sherpa, then I have used 3 separate model components to create my model. These include the power law (pl), the lorentzian (l1) and the gaussian (g1). The power law describes the continuum, and the three together represent the data.
To measure the equivalent width of the K line, I use Sherpa’s eqwidth() function. The syntax is:
>>>eqwidth(continuum, continuum+line, lo=lower wl limit, hi=upper wl limit, id=data ID #)
Using my data for a K line whose line goes from 1.252µ to 1.253µ, I measure the eq. width like so:
>>> eqwidth(pl, pl+l1+g1, lo=1.252, hi=1.253, id=1)
The output will be the equivalent width value, in microns. Nice and simple. The hard part was constructing the models and eyeballing the lower/upper limits for the eq. width measurement. All of this I did manually by eye.
The line-to-continuum flux ratio and full width half maximum measurements are harder. They require locating the actual arrays of values for the various models. In order to access the model’s x and y values (continuum+lorentzian+gaussian), you do:
>>> modelArrays = get_model_plot(1) # The 1 is the ID number
# modelArrays.x –> array of x values
# modelArrays.y –> array of y values
To access the y values for the continuum only (pl), you have to do:
>>> key = pl._cache.keys()[0]
>>> cont_yvals = pl._cache[key]
# This may not work if some of the values in pl.__dict__ are not set properly. To be honest, # I’m not fully sure which ones may screw this up, but for me, I have:
>>> pl._NoNewAttributesAfterInit__initialized = True
>>> pl._use_caching = True
# For me, since I use a power law with an exponent of 0, all of the y values are the same. So I can access this value instead by doing:
>>> cont_yval = pl.ampl # The continuum is a constant value: the amplitude
Armed with the model’s x and y value arrays, and the continuum’s y value array (or value), we can make the measurements. To calculate the line-to-continuum flux ratio, we must first find the minimum (maximum for emission lines) value of the model, and then compare it with the corresponding continuum flux value:
>>> F_model = numpy.min(modelArrays.y) # Minimum of the model
>>> min_ind = numpy.where(modelArrays.y == F_model)[0][0] # min’s array index
>>> F_cont = cont_yvals[min_ind] # Continuum value
>>> depth = 1.0 – (F_model/F_cont) # The line-to-continuum flux ratio!
The full width half maximum was a little tougher. The algorithm I used is as follows:
>>> F_halfmax = (F_cont + F_max) / 2.0 # Flux at the half maximum
>>> indices = numpy.where(modelArrays.y <= F_halfmax)[0]
>>> wls = modelArrays.x[indices]
>>> fwhm = wls[-1] – wls[0]
The problem with this algorithm is that sometimes there are pockets of spectrum that are equal to or lower than the half max, which get into the wls array. For those, I had to go through them manually and fix the wls array.
That’s how I measured the equivalent width, line-to-continuum flux ratio, and full width half maximum using Sherpa. If you have any questions or know of a better (or different) way of calculating these values, feel free to leave a comment or email me at danfeldman90@gmail.com